#include <alignment_scorer.hpp>
Public Member Functions | |
| virtual | ~AlignmentScorer ()=default |
| virtual int32_t | score_alignment (const Alignment &aln) const =0 |
| Compute a single integer score for the alignment under this scorer's scheme. More... | |
| virtual double | get_log_base () const =0 |
Static Protected Member Functions | |
| static double | recover_log_base (const double matrix[16], double gc_content, double tol=1e-12) |
Static Private Member Functions | |
| static bool | verify_valid_log_odds_score_matrix (const double matrix[16], const double nt_freqs[4]) |
| Check a score matrix to make sure it has negative scores for random sequence. More... | |
| static double | alignment_score_partition_function (double lambda, const double matrix[16], const double nt_freqs[4]) |
Detailed Description
Base interface for computing scores for alignments under some (not necessarily affine-gap) scoring scheme.
Constructor & Destructor Documentation
◆ ~AlignmentScorer()
|
virtualdefault |
Member Function Documentation
◆ alignment_score_partition_function()
|
staticprivate |
Partition function used for recovering the log base for a scoring scheme. TODO: What is a partition function exactly?
◆ get_log_base()
|
pure virtual |
Get the log base value used to probabilistically interpret scores. Implementations probably need to have a stores GC content to implement this.
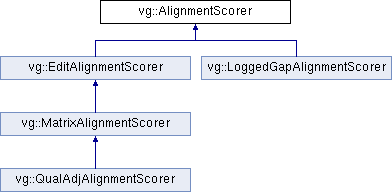
Implemented in vg::LoggedGapAlignmentScorer, and vg::MatrixAlignmentScorer.
◆ recover_log_base()
|
staticprotected |
Bisects to find the log base under which the partition function over the given 4x4 substitution matrix and nucleotide frequencies (from GC content) equals 1.
Useful for implementing the recover_log_base operation for a derived class, if that class can produce a scoring matrix.
◆ score_alignment()
|
pure virtual |
Compute a single integer score for the alignment under this scorer's scheme.
Implemented in vg::LoggedGapAlignmentScorer, and vg::EditAlignmentScorer.
◆ verify_valid_log_odds_score_matrix()
|
staticprivate |
Check a score matrix to make sure it has negative scores for random sequence.
The documentation for this class was generated from the following files:
 1.9.1
1.9.1