|
| | GSSWAligner (GSSWAligner &&)=default |
| |
| GSSWAligner & | operator= (GSSWAligner &&)=default |
| |
| | GSSWAligner (const GSSWAligner &)=delete |
| |
| GSSWAligner & | operator= (const GSSWAligner &)=delete |
| |
| virtual void | align_pinned (Alignment &alignment, const HandleGraph &g, bool pin_left, bool xdrop=false, uint16_t xdrop_max_gap_length=default_xdrop_max_gap_length) const =0 |
| |
| virtual void | align_pinned_multi (Alignment &alignment, vector< Alignment > &alt_alignments, const HandleGraph &g, bool pin_left, int32_t max_alt_alns) const =0 |
| |
| virtual void | align_global_banded (Alignment &alignment, const HandleGraph &g, int32_t band_padding=0, bool permissive_banding=true, uint64_t max_cells=std::numeric_limits< uint64_t >::max()) const =0 |
| |
| virtual void | align_global_banded_multi (Alignment &alignment, vector< Alignment > &alt_alignments, const HandleGraph &g, int32_t max_alt_alns, int32_t band_padding=0, bool permissive_banding=true, uint64_t max_cells=std::numeric_limits< uint64_t >::max()) const =0 |
| |
| virtual void | align_xdrop (Alignment &alignment, const HandleGraph &g, const vector< MaximalExactMatch > &mems, bool reverse_complemented, uint16_t max_gap_length=default_xdrop_max_gap_length) const =0 |
| | xdrop aligner More...
|
| |
| virtual void | align_xdrop (Alignment &alignment, const HandleGraph &g, const vector< handle_t > &order, const vector< MaximalExactMatch > &mems, bool reverse_complemented, uint16_t max_gap_length=default_xdrop_max_gap_length) const =0 |
| |
| virtual void | align (Alignment &alignment, const HandleGraph &g, bool traceback_aln) const =0 |
| |
The basic GSSW-based core aligner implementation, which can then be quality-adjusted or not. Owns its scorer and MAPQ calculator; all scoring and MAPQ work goes through those, not through the aligner.
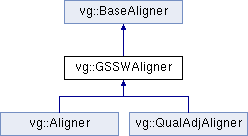
You aren't meant to instantiate this directly; you should have an Aligner or a QualAdjAligner as the concrete class instead.
| virtual void vg::GSSWAligner::align_global_banded_multi |
( |
Alignment & |
alignment, |
|
|
vector< Alignment > & |
alt_alignments, |
|
|
const HandleGraph & |
g, |
|
|
int32_t |
max_alt_alns, |
|
|
int32_t |
band_padding = 0, |
|
|
bool |
permissive_banding = true, |
|
|
uint64_t |
max_cells = std::numeric_limits< uint64_t >::max() |
|
) |
| const |
|
pure virtual |
Store top scoring global alignments in the vector in descending score order up to a maximum number of alternate alignments (including the optimal alignment). If there are fewer than the maximum number of alignments in the return value, then the vector contains all possible alignments. The optimal alignment will be stored in both the vector and the original alignment object.
When multiple alignments have the same score, they are ordered deterministically but arbitrarily.
Throws BandMatricesTooBigException if the max_cells limit on DP matric size is hit.
Implemented in vg::QualAdjAligner, and vg::Aligner.
| virtual void vg::GSSWAligner::align_pinned |
( |
Alignment & |
alignment, |
|
|
const HandleGraph & |
g, |
|
|
bool |
pin_left, |
|
|
bool |
xdrop = false, |
|
|
uint16_t |
xdrop_max_gap_length = default_xdrop_max_gap_length |
|
) |
| const |
|
pure virtual |
store optimal alignment against a graph in the Alignment object with one end of the sequence guaranteed to align to a source/sink node. if xdrop is selected, use the xdrop heuristic, which does not guarantee an optimal alignment.
pinning left means that that the alignment starts with the first base of the read sequence and the first base of a source node sequence, pinning right means that the alignment starts with the final base of the read sequence and the final base of a sink node sequence
Gives the full length bonus only on the non-pinned end of the alignment.
Implemented in vg::QualAdjAligner, and vg::Aligner.
 Public Member Functions inherited from vg::BaseAligner
Public Member Functions inherited from vg::BaseAligner 1.9.1
1.9.1